Get Involved
We believe understanding the microbiome is essential to prevent and treat diseases that are rising worldwide — and that collaborating across institutions and sectors is key to accelerating progress.
We seek to connect people, techniques, and technologies to accelerate the advent of microbiome-based approaches to improve public health.
If you would like to collaborate, sponsor a research project, or connect to our network of scientists, engineers, and clinicians, please email humanmicrobiome@mit.edu or fill out a contact form through one of the links below.
If you would like to learn more about microbiome seminars and events at MIT, you can find out more and get involved through the MIT Microbiome Club.
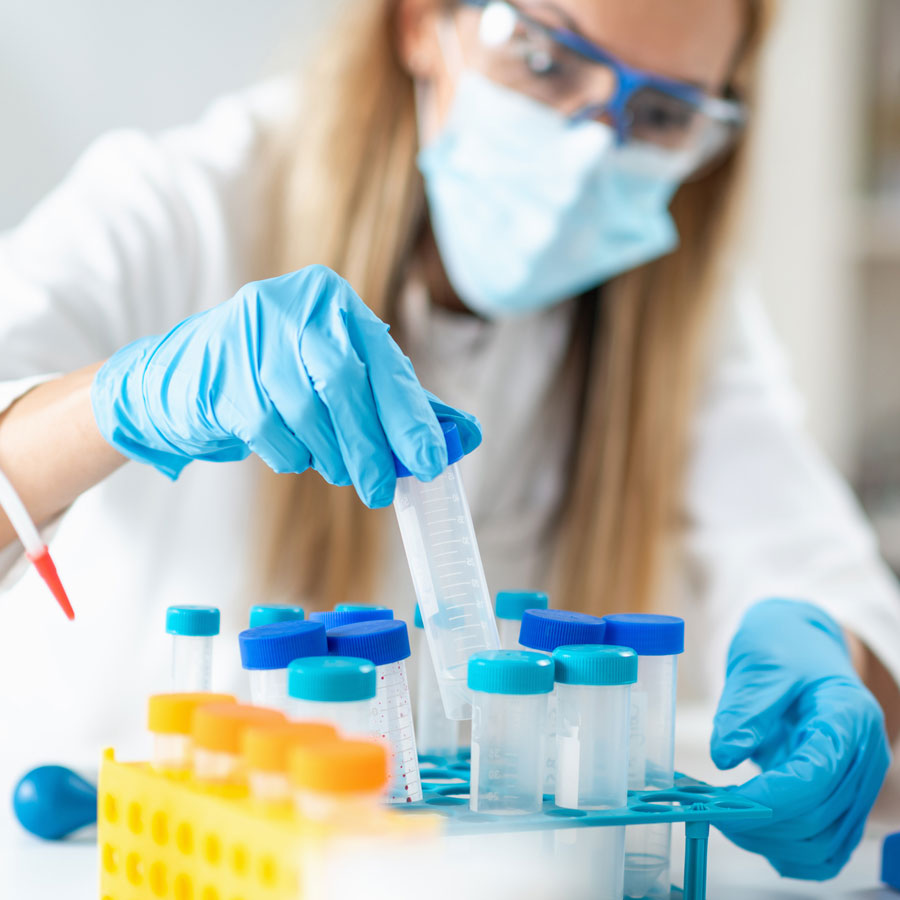
For Researchers
To develop the spectrum of different therapies needed for patients with microbiome-associated disease, CMIT needs to attract the best minds to join our community.
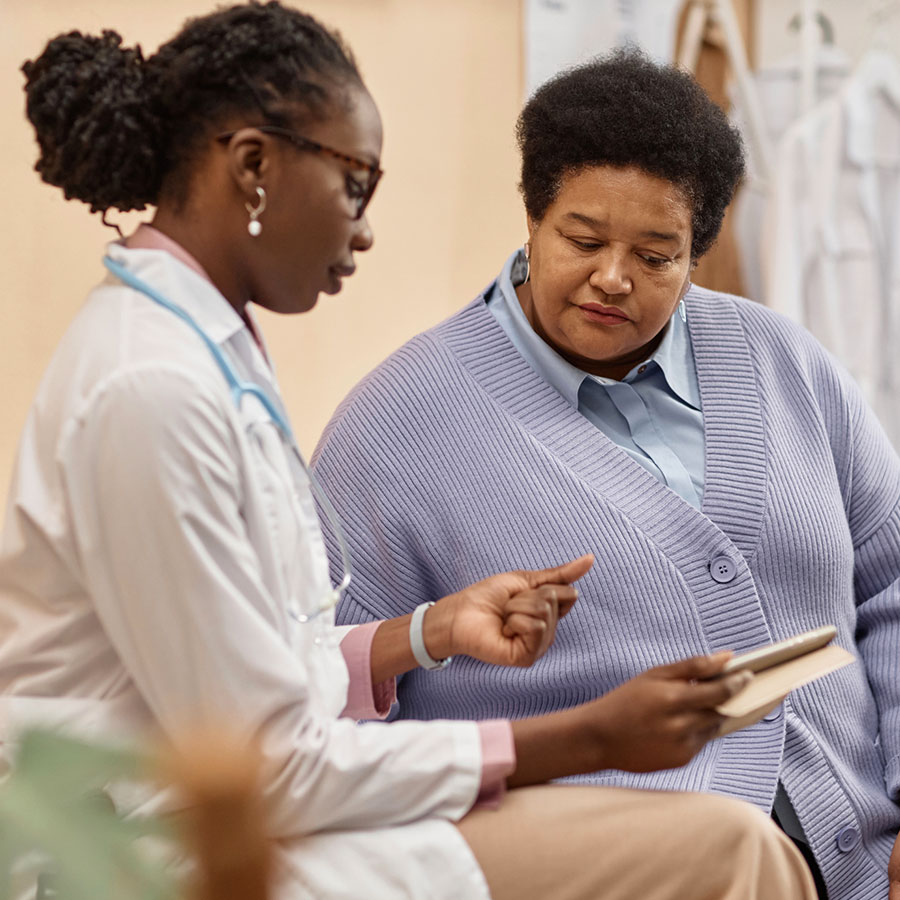
For Clinicians
CMIT focuses on human observations of the gut microbiome. Our clinical partnerships are essential both to understand the microbiome in patients with microbiome-associated disease, and to translate our research into the clinical setting.
For Industry
CMIT focuses on human observations of the gut microbiome. Our clinical partnerships are essential both to understand the microbiome in patients with microbiome-associated disease, and to translate our research into the clinical setting.
For Students & Postdocs
CMIT is committed to educating the next generation of researchers, physicians, industry leaders, technologists, and policy makers on the importance of the microbiome for human health.
Mailing Lists
Sign up for email updates on CMIT Seminar and Events or the Microbiome Club.
Donate
You are committed to a better future in healthcare. CMIT wants to make it a reality. Help us realize low-cost, safe, and effective treatments for today’s biggest challenges in healthcare. Partner with us on the path to better health.