Cranking up the resolution
by Lindzi Wessel
New Science paper presents cutting-edge method for examining the microbiome
Technical limitations have long challenged our ability to map genomes across a microbiome. CMIT researchers and their collaborators have figured out how to circumvent those obstacles and enhance our understanding of the invisible communities in our guts.
No two gut microbiomes are the same. A unique assembly of bacteria and other tiny lifeforms resides in each of our digestive systems, helping to determine what foods we can tolerate, how we absorb nutrients and what ailments we might be prone to. Characterizing these invisible communities relies on our ability to decipher the genomes of each member, but technical constraints in how we gather microbial DNA have long limited scientists’ ability to conduct such surveys.
Now, researchers from MIT’s Center for Microbiome Informatics and Therapeutics and Harvard University have presented a new approach to analyzing the gut microbiome, allowing them to catalog nearly a hundred species per microbiome sample, more than previously available methods have allowed. The technique, dubbed MicrobeSeq and detailed in a Science paper published earlier this month, will enable researchers to study any microbiome with more precision and speed than ever before, the paper’s authors say, opening the door to a better understanding of how our individualized microbes affect our health and risk of disease.
“This is a new high-throughput survey of a human gut microbiome sample,” says Shijie “Jay” Zhao, a former MIT graduate student and co-first author of the new study. “We’re now a lot closer to being able to determine what each and every member of a microbiome looks like.”
A troublesome tradeoff
For decades, characterizing a microbe’s DNA required an arduous process of growing that microbe in the lab—coaxing it to replicate again and again in a dish until scientists could collect enough genetic material to perform their analyses. This process is resource and time consuming, making it impractical when trying to understand the behavior and evolution of hundreds of microbial species interacting within a community. The advent of so-called metagenomic sequencing, however, means scientists can simply pluck all the genetic material out of a sample and quickly read out all the DNA within, letting them examine hundreds of genomes at the same time.
But, though fast and efficient, metagenomics has its drawbacks, particularly when applied to microbes. Before DNA can be read out by a sequencer, it must be collected. That means popping the walls or membranes that enclose the cells in a sample and letting their DNA spill out for collecting. Then, using a particular portion of DNA unique to each species—a natural barcode of sorts—researchers can read out DNA from all the species present within their sample. This methodology would work perfectly if the genomes of each member of a microbial species were the same. But because of how microbes reproduce and exchange DNA, the genomes of two microbes of a single species can vary widely from one another. This means that, though metagenomics lets researchers quickly capture a large breadth of information about a microbial community, it forces them to sacrifice subtle detail that may have major consequences for human health
For example, even though two strains of E. coli, would share the same species barcode, they might have less than 50 percent of their DNA in common, says Eric Alm, Co-Director of CMIT and an MIT professor of Biological Engineering. The large genetic differences can translate to substantial changes in how the microbes function. This helps explain why some strains of E. coli coexist harmoniously with the human gut while others can cause dangerous afflictions of vomiting and diarrhea.
“It’s fairly shocking how extreme these differences can be,” says Alm, who shares senior authorship of the new paper. “And most studies out there are not going to be able to tell you what strain you have, which may be one reason why, when we study the microbiome, things aren’t as clear as we’d like them to be.”
Using just a snippet of DNA as a species barcode, then, is not enough to characterize a whole population of microbes, which—to complicate things further—sometimes spit out chunks of DNA for neighboring microbial cells to pick up. When DNA from all the cells of a microbiome are broken into chunks and mixed together, it’s impossible to tell which chunks of DNA came from which cells. Current technology has thus forced researchers to choose between sacrificing breadth—in the case of single cell culture methods, or to sacrifice precision when using metagenomics. Alm and his team, however, wanted a method that would let them have both.
Your microbiome in tiny little droplets
The solution came through microfluidics. Alm’s team collaborated with David Weitz, a co-senior author of the new paper and a professor of Harvard University who co-invented a high throughput RNA-seq method for sequencing individual cells using microfluidics. Co-first author Wenshan Zheng on Weitz’s team worked closely with researchers in the Alm lab to develop a series of tiny fluid-filled reactors that, together, prepare microbial DNA for sequencing without losing track of which DNA came from which microbe.
First, in a reactor chip no bigger than a credit card, microbial cells are separated and each is encapsulated inside microscopic fluid droplets, separate from other microbes. Here, the cell is broken open, but instead of being released into one big pool of genetic material, each individual microbe’s DNA remains captured inside its corresponding droplet. In two additional microfluidic devices, the DNA is then prepped for processing without leaving the droplets. The fourth and last microfluidic device is key: Here each droplet containing microbial DNA merges with other droplets containing artificial DNA barcodes manufactured by the researchers in the lab. Now, instead of relying on the naturally occurring species barcode only present in one snippet of DNA, researchers can label the entire genome with a barcode based on the droplet—and thus the unique microbe—that DNA came from.
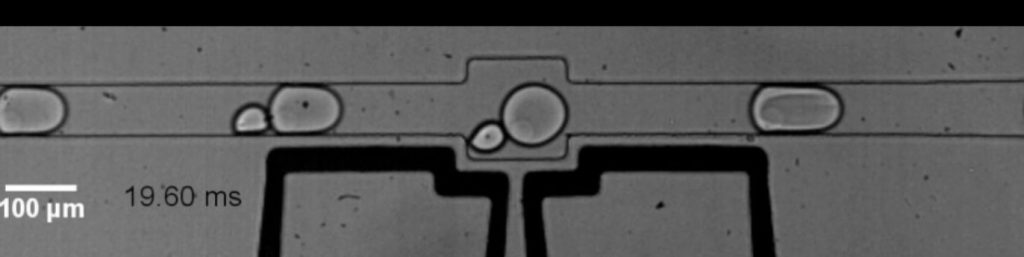

To validate their new method, the team used MicrobeSeq to read around 20,000 microbes collected from the microbiome of a previously studied research participant and stitched the reads into genome maps for 76 bacterial species. The MicrobeSeq results closely matched previous findings the team got from this microbiome using laborious, gold-standard culture methods.
Eventually, the researchers say, MicrobeSeq could be used to help link certain microbial strains to disease or food intolerances and to monitor changes in an individual’s microbiome over time. The technique can also be used to study how bacteria-targeting viruses might influence our guts. In their paper, the researchers reported that MicrobeSeq indeed picked up the DNA of viruses attached to gut microbes and indicated which strains of microbes were being targeted. And the precision afforded by the technique is not limited to studying the gut. It could theoretically be used in any microbial population, including marine or soil ecosystems.
“MicrobeSeq opens up this future where we can look at a microbiome and put together a list of all the species that are there together with a list of all the genes that are in the species,” says Alm. “In the case of the gut, it means we can really start to understand how different microbial genes are contributing to real health effects.”
Other MIT-affiliated authors were Yehang Yin, David M. Needham, Ethan D. Evans and Chengzhen L. Dai. Additional authors were Harvard University researchers Huidan Zhang and co-senior author Peter J. Lu.